Aims and scope
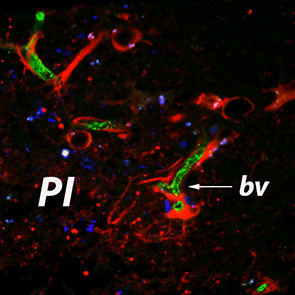
Vascular Cell represents an open access medical journal which concentrates on publication of a wide range of topics related to the vascular system including neo-vascularization and angiogenesis. Since vascular endothelial cells represent a crucial factor in numerous pathological conditions such as: cancer, stroke, myocardial infarction or atherosclerosis their behavior is of paramount importance for the clinical outcomes of a wide range of patients.
VascularCell indexing
We are in the process of reindexing VascularCell with PubMed. Currently PubMed holds the archive of Vascular Cell articles published between 2009-2017
Latest Articles
VascularCell articles from Volume 15 (issue 1), published in 2025.